Nakagawa Lab
Principal Investigator
In the Nakagawa Lab, our work begins with a pressing clinical reality: esophageal squamous cell carcinoma (ESCC) remains one of the most lethal cancers worldwide, with survival rates that have changed far too little despite decades of effort. Our research is motivated by the belief that better treatments will only emerge from a deeper understanding of how these cancers arise, evolve, and resist therapy. For more than thirty years, the lab has followed the biology of the upper aerodigestive epithelium from normal tissue to precancerous change to aggressive malignancy—connecting fundamental discoveries in cell signaling, tumor heterogeneity, and environmental risk factors to real clinical need. Today, the backbone of our work is a powerful patient-derived organoid platform that allows us to recreate the spectrum of human disease in the lab with unprecedented fidelity, while capturing patient disease heterogeneity. These living models enable us to capture the complexity of tumors, study how DNA damage and tissue stress drive cancer initiation, and test thousands of potential therapeutics in a physiologically relevant context. By combining organoid engineering, genetic manipulation, and large-scale drug screening, we aim to bridge the gap between mechanism and medicine—transforming biological insight into new, therapeutic strategies for patients facing ESCC and other upper aerodigestive cancers.
Research Objectives
Decoding the origins of ESCC: Modeling carcinogenesis and therapeutic vulnerability

Patient-derived organoid platforms capture ESCC evolution and enable high-throughput discovery of clinically actionable therapeutic vulnerabilities.
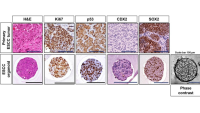
Our lab investigates how esophageal squamous cell carcinoma (ESCC) arises and evolves, with a focus on the molecular events that transform normal epithelium into therapy-resistant cancer. Using patient- and mouse-derived organoids, engineered cell systems, and complementary in vivo models, we recreate the continuum from normal tissue to preneoplasia to invasive carcinoma and even metastasis. These platforms allow us to probe oncogenic signaling pathways, cellular heterogeneity, and evolutionary pressures that shape tumor behavior. Coupled with high-throughput screening of large compound libraries, our work identifies vulnerabilities that can be translated into more effective treatments.
This research establishes a mechanistic foundation for precision therapy in ESCC, enabling drug discovery efforts in models that closely reflect patient biology. These models provide the experimental backbone that supports virtually all other research objectives in the lab, including environmental exposures, DNA damage responses, and epithelial stress pathways.
Environmental risk, metabolism, and DNA damage in esophageal pathobiology

Models of esophageal epithelial progression (normal → preneoplasia → SCC) provide a platform to define how alcohol exposure and impaired ALDH2-mediated metabolism drive cellular stress, DNA damage, and carcinogenesis.
Alcohol exposure remains one of the strongest environmental risk factors for ESCC, yet the cellular mechanisms linking exposure to cancer progression are far from completely understood. This objective explores how metabolic stress and impaired mitochondrial homeostasis influence epithelial cell fate and carcinogenesis, particularly in the context of ALDH2 biology, responsible for alcohol detoxification in the cell. Through organoid systems, genetically modified models, and mechanistic molecular studies, we examine how environmental insults reshape tissue responses, promote genomic instability, and create conditions permissive for cancer initiation.
By defining how common exposures drive disease at the cellular level, this work opens pathways for prevention strategies and targeted interventions aimed at reducing cancer risk. Findings from this objective directly inform our ESCC models and connect closely to studies of DNA repair pathways, including Fanconi anemia–associated vulnerability.
DNA repair deficiency and stem cell vulnerability in squamous carcinogenesis

Disruption of FA-mediated DNA repair in organoid systems reveals how environmental stress induces genome instability, reshapes epithelial cell survival, and creates vulnerabilities that drive SCC pathogenesis.
Inherited defects in DNA repair pathways, such as those seen in Fanconi anemia (FA), create profound susceptibility to young-onset squamous cell carcinomas (SCC). This objective uses organoid models to investigate how epithelial cells with impaired DNA damage responses react to environmental stress and accumulate oncogenic alterations. By modeling these interactions in physiologically relevant systems, we aim to identify early events that predispose cells to malignant transformation as well as DNA repair pathway addiction of SCC.
This work provides critical insight into cancer prevention in high-risk populations while uncovering universal mechanisms of DNA repair failure and addiction relevant to sporadic SCC. Mechanisms uncovered here feed directly into our broader studies of environmental risk and SCC pathogenesis, informing both mechanistic understanding and therapeutic targeting.
Tissue stress, autophagy, and epithelial remodeling
Epithelial tissues constantly respond to inflammatory and environmental stress, and these responses can shape long-term disease risk. This objective investigates how autophagy and stress-response pathways regulate epithelial remodeling in esophageal disease states. Using advanced 3D culture systems and molecular analysis, we examine how adaptive cellular processes influence tissue architecture, survival, and progression toward pathology in the context of Eosinophilic Esophagitis, a chronic inflammatory disease of the esophagus, driven by food allergens.
Understanding how epithelial tissues adapt—or fail to adapt—to stress reveals mechanisms that may underlie both inflammatory disease and malignant transformation. Insights into autophagy and remodeling provide important biological context for our cancer-focused projects, highlighting shared pathways between tissue repair, chronic inflammation, and carcinogenesis.
Building translational infrastructure: Organoid innovation and shared scientific platforms
(P30DK132710 — Organoid and Cell Culture Core, OCCC)
A central mission of the lab is the development and dissemination of cutting-edge three-dimensional culture systems that bring human biology into experimental research. Through leadership of the Organoid and Cell Culture Core (OCCC), we develop, engineer, and distribute well-annotated organoid models derived from patients and mouse models, spanning normal tissue, preneoplasia, and cancer. The core also provides training, genetic engineering support, and scalable platforms for collaborative research across the cancer and digestive disease communities.
By enabling investigators to work in physiologically relevant models, the OCCC accelerates discovery, improves reproducibility, and supports translational precision oncology research at scale. The organoid platform is the unifying axis across all research objectives, allowing discoveries in one project to rapidly inform and advance others.
To learn more about the OCCC, please visit us at our DLDRC webpage here.
Together, these objectives form an integrated research ecosystem in which organoid-based modeling bridges basic mechanisms, environmental risk, and therapeutic discovery — accelerating the translation of biologic insight into improved outcomes for patients with esophageal and upper aerodigestive cancers.
Lab Members
Sam Flashner, PhD
- Associate Research Scientist

Kento Kawasaki, MD, PhD
- Postdoctoral Research Scientist

Debora Cavero Moreno, PhD
- Postdoctoral Research Scientist

Jiayue (Wendy) Wu
- PhD Student

Cecilia Martin, MS
- Lab Manager, Nakagawa Lab
- Technical Director, OCCC

Select Publications
Morimoto M, Kawasaki K, McNamee N, Flashner S, Shimonosono R, Shimonosono M, Matsuura N, Tomita Y, Hirose W, Teranishi R, Itami T, Guha M, Rajagopalan P, Martin C, Golden H, Dhakal D, Wilkins BJ, Klein-Szanto AJ, Wangensteen KJ, Abrams JA, Pomenti S, Katzka DA, Que J, Whelan KA, Muir AB, Kita H, Wright BL, Doyle AD, Nakagawa H, Sachdeva UM. Mitochondrial dysfunction drives basal cell hyperplasia in eosinophilic oesophagitis. Gut. 2025 Sep 8;74(10):1571-1588. doi: 10.1136/gutjnl-2024-334561. PMID: 40701793; PMCID: PMC12438975.
Shimonosono M, Morimoto M, Hirose W, Tomita Y, Matsuura N, Flashner S, Ebadi MS, Okayasu EH, Lee CY, Britton WR, Martin C, Wuertz BR, Parikh AS, Sachdeva UM, Ondrey FG, Atigadda VR, Elmets CA, Abrams JA, Muir AB, Klein-Szanto AJ, Weinberg KI, Momen-Heravi F, Nakagawa H. Modeling Epithelial Homeostasis and Perturbation in Three-Dimensional Human Esophageal Organoids. Biomolecules. 2024 Sep 5;14(9):1126. doi: 10.3390/biom14091126. PMID: 39334892; PMCID: PMC11430971.
Flashner S, Shimonosono M, Tomita Y, Matsuura N, Ohashi S, Muto M, Klein-Szanto AJ, Alan Diehl J, Chen CH, Mochly-Rosen D, Weinberg KI, Nakagawa H. ALDH2 dysfunction and alcohol cooperate in cancer stem cell enrichment. Carcinogenesis. 2024 Feb 12;45(1-2):95-106. doi: 10.1093/carcin/bgad085. PMID: 37978873; PMCID: PMC10859731.
Parikh AS, Yu VX, Flashner S, Okolo OB, Lu C, Henick BS, Momen-Heravi F, Puram SV, Teknos T, Pan Q, Nakagawa H. Patient-derived three-dimensional culture techniques model tumor heterogeneity in head and neck cancer. Oral Oncol. 2023 Mar;138:106330. doi: 10.1016/j.oraloncology.2023.106330. Epub 2023 Feb 9. PMID: 36773387; PMCID: PMC10126876.
Flashner S, Martin C, Matsuura N, Shimonosono M, Tomita Y, Morimoto M, Okolo O, Yu VX, Parikh AS, Klein-Szanto AJP, Yan K, Gabre JT, Lu C, Momen-Heravi F, Rustgi AK, Nakagawa H. Modeling Oral-Esophageal Squamous Cell Carcinoma in 3D Organoids. J Vis Exp. 2022 Dec 23;(190):10.3791/64676. doi: 10.3791/64676. PMID: 36622034; PMCID: PMC10037110.
Flashner S, Yan KS, Nakagawa H. 3D Organoids: An Untapped Platform for Studying Host-Microbiome Interactions in Esophageal Cancers. Microorganisms. 2021 Oct 20;9(11):2182. doi: 10.3390/microorganisms9112182. PMID: 34835308; PMCID: PMC8622040.
Karakasheva TA, Kijima T, Shimonosono M, Maekawa H, Sahu V, Gabre JT, Cruz-Acuña R, Giroux V, Sangwan V, Whelan KA, Natsugoe S, Yoon AJ, Philipone E, Klein-Szanto AJ, Ginsberg GG, Falk GW, Abrams JA, Que J, Basu D, Ferri L, Diehl JA, Bass AJ, Wang TC, Rustgi AK, Nakagawa H. (2020). Generation and Characterization of Patient-Derived Head and Neck, Oral, and Esophageal Cancer Organoids. Current Protocols in Stem Cell Biology. 53(1):e109. PMCID: PMC7350550.






